Research
Our biological interests span a broad range of different research topics around the common theme of cellular organization and dynamics. Our groups investigate fundamental biological questions in areas such as chromatin organization and gene expression, intracellular compartmentalization and transport, cell growth, division and homeostasis, as well as dynamic transitions involved in these processes at different scales. We seek to understand how the dynamic interplay of biomolecules, especially of proteins, nucleic acids, and lipids forms the fundamental functional unit of life – a cell. To address our biological questions, we use interdisciplinary approaches and exploit state-of-the-art technologies, first and foremost advanced microscopy.
The research groups located at the Institute of Biochemistry exhibit a high degree of interaction and integration. This is apparent in our joint seminars and the ongoing collaborations within the institute. In addition, we have a common management for our funding, human resources and a well-established IT infrastructure.
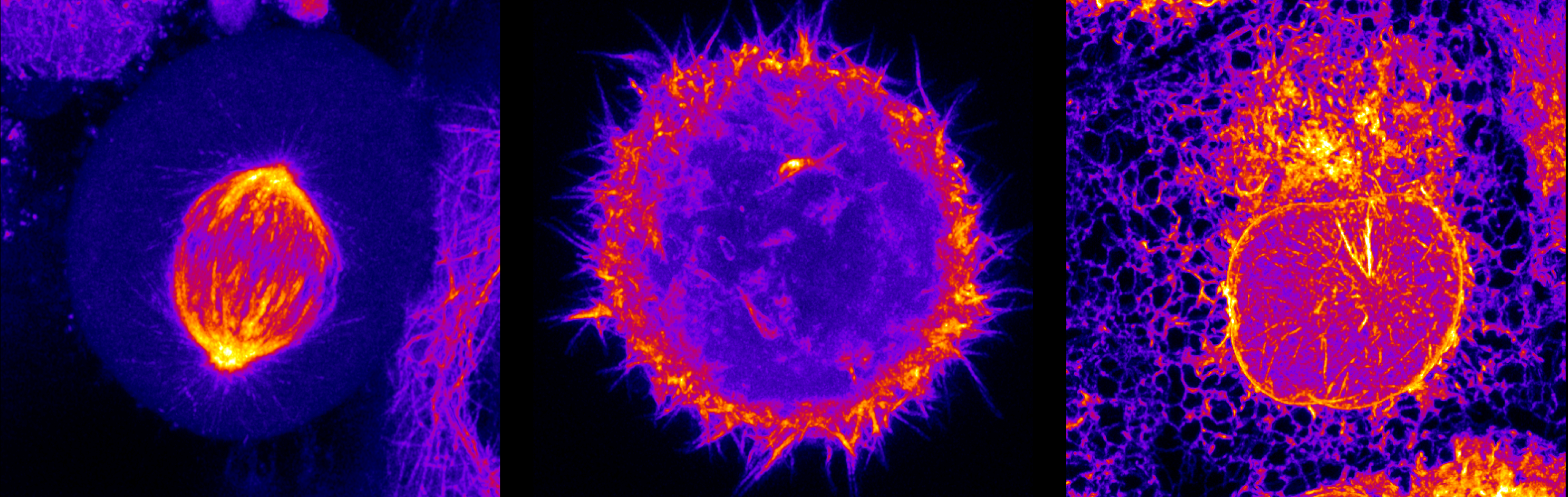
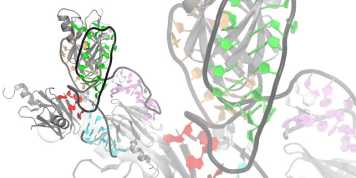
We are interested in understanding at atomic level the interaction between proteins and RNA at the heart of pre-mRNA splicing and/or RNA editing.
Our focus is on determining the structures of protein-RNA complexes using Nuclear Magnetic Resonance (NMR) in solution. In addition we also employ other biophysical and biochemical methods to gain further insights.
Cells are the basic unit of life and have acquired in evolution not only the ability to proliferate but also to take decisions, protect themselves against a broad variety of stresses or pathogens, memorize information about their past experience and adapt their shape and activities to environmental cues and conditions. We are using different yeast species to study how Eukaryotes carry out these different functions and how these activities contribute to cellular diversity, speciation and aging. We focus particularly on the role of two universal processes in Eukaryotes, namely cell division (particularly asymmetric cell division) and the sexual cycle (focusing on mating). More...
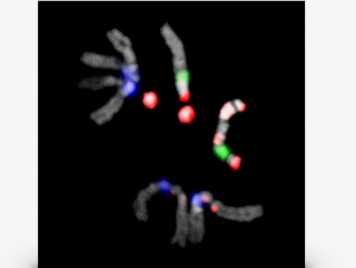
We are broadly interested in exploring the function(s) of satellite DNA repeats at a cellular and organismal level and in determining whether the widely observed divergence of satellite DNAs is one of the fundamental barriers that reproductively isolates different species. More...
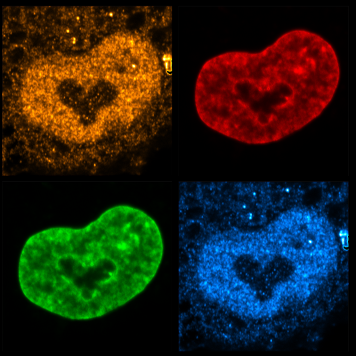
Our laboratory is interested in the organization, function and dynamics of the human cell nucleus. The nucleus is a truly fascinating organelle that is perfectly adapted to its functions as cellular command center and protective home of our genome. We are investigating different aspects of nuclear biology, including the fundamental biosynthetic function of the nucleus in the assembly of ribosomal subunits, the role of the nuclear envelope in nucleo-cytoplasmic communication and genome organization as well as the dynamics of the nuclear compartment during the cell cycle. By our research, we seek to improve our understanding of how nuclear dysfunction contributes to human diseases such as ribosomopathies, nuclear envelopathies and cancer. More...
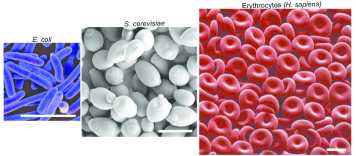
From bacteria to higher eukaryotes all cells of a specific cell type have a very characteristic size. Why tight regulation of cell size is so critical for cell function is largely unexplored. This question is interesting for two reasons: Deviation from normal cell size occurs during non-physiological conditions such as during cell senescence and in tumors and coincides with altered cell function, strongly suggesting that having a specific cell size is critical for cell function. On the other hand, cells in multicellular organisms exist at sizes that span several orders of magnitude, but every cell type has a well-defined size. This raises the possibility that cell size may contribute to defining cell type specific cell function. In our lab we use a combination of genetic, live cell microscopy and molecular biology approaches to understand how cell size affects different aspects of cell physiology.
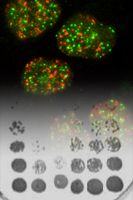
Regulation of cell growth and division by selective degradation mechanisms. Every living organism requires a tight coordination of cell growth and cell division, and defects in these regulatory systems are intimately linked to metabolic disorders and cellular transformation. Cell growth is manifested by an increase in cell mass, and thus involves the homeostasis and regulation of metabolic pathways, ribosome biogenesis and protein translation, and catabolic pathways including autophagy and ubiquitin-dependent degradation. Cell division requires the precise duplication of chromosomes and their partitioning between two daughter cells. The morphological transitions that lead to chromosome segregation during mitosis need to be coordinated in space and time with subsequent cytoplasm separation during cytokinesis. However, the molecular mechanisms that govern cell growth and division, and the different intra- and extracellular signals that regulate these pathways, are still poorly understood. More...
The interest of our laboratory is centered on very fundamental questions in eukaryotic cell biology: how are macromolecules properly localized within a cell, and how do eukaryotes take advantage of transport and compartmentalization to regulate their gene expression programs? Our work particularly focuses on studying transport processes between the nucleus and the cytoplasm. The exchange of macromolecules between the nucleus and the cytoplasm is absolutely essential to establish and maintain order in eukaryotic cells, and constitutes an important step in the regulation of gene expression. Our laboratory combines genetic, biochemical, cell biological and biophysical approaches in Saccharomyces cerevisiae and in metazoan cells to characterize and analyze the molecular machinery that is responsible for the transport of macromolecules into and out of the nucleus.
More...